Unprecedented Tracking of Chromatin Dynamics Shows How Genome Stability Emerges

A new study reveals genome motion with unprecedented temporal resolution, offering insight into how cells maintain genome stability. Chromatin, the DNA-protein complex that organizes our genome, is constantly moving inside the nucleus. Although constrained by its chain‑like structure, this motion determines how efficiently distant genomic regions find one another, from enhancers activating genes to broken DNA ends reuniting after damage.
For years, limited measurement timescales have left chromatin dynamics poorly understood. Anders Sejr Hansen and his team, including Matteo Mazzocca, Domenic Narducci, and Simon Grosse Holz, overcame this barrier by combining MINFLUX microscopy with camera-based tracking, capturing chromatin motion across seven orders of magnitude in time.
They identified two major motion regimes: some cell types keep chromatin tightly confined, while others allow broader exploration over longer timescales.
These dynamics help explain how cells maintain genome stability and influence the mechanisms that govern DNA repair and gene regulation. Strong confinement during DNA double‑strand breaks helps broken ends stay close, supporting efficient repair despite the 10–50 breaks human cells experience daily. In gene regulation, nearby enhancers (<100 kb) can reach their target genes by passive diffusion, while distant enhancers (>1 Mb) likely rely on active, energy‑driven mechanisms that enable long‑range communication. Because such distant enhancer–gene interactions often activate oncogenes in cancer, disrupting these active mechanisms could offer a therapeutic strategy.
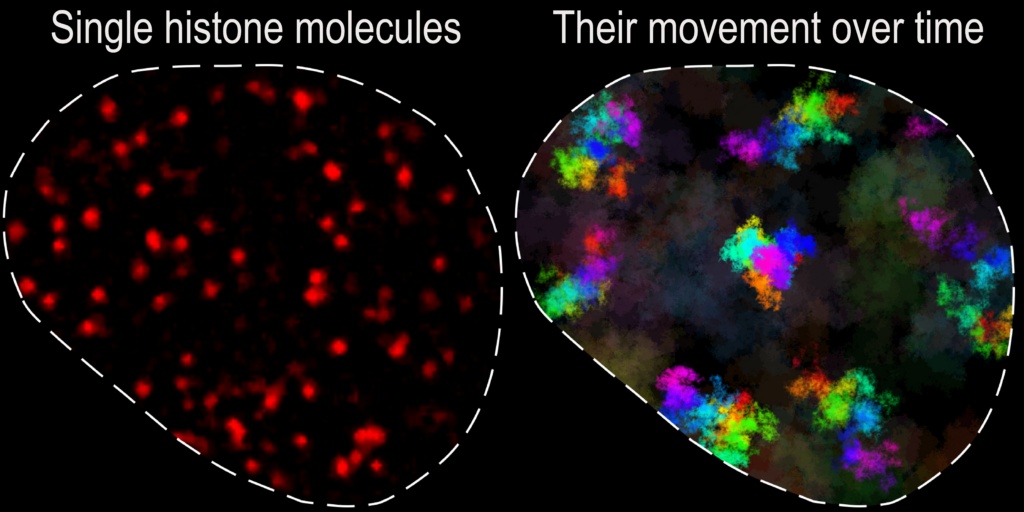
Left: a snapshot image of individual locations (histone proteins) of the genome visualized inside the cell nucleus. Right: Reconstruction of the “movement territories” of individual loci of the genome.
Read the full article here. Paid subscription to Nature Structural & Molecular Biology required.