A hidden mechanism for DNA synthesis

Bacteria possess a vast array of molecular strategies to fight off viral infections. While well-known defenses like CRISPR have become essential tools for biotechnology, they represent only a fraction of the landscape, and many other defense systems remain uncharacterized.
In a study published in Science, Alex Gao and his team identified a defense system called DRT3 that builds a DNA double helix in a previously unknown way. This system constructs a specific, repeating DNA sequence using two distinct enzymes known as reverse transcriptases.
Normally, cells build DNA by reading an existing genetic blueprint. One enzyme in this system follows this familiar pathway. But in a surprising twist, the second builds its strand without a template, assembling a precise matching sequence from scratch to complete the double helix.
This finding reveals an unexpected level of biochemical ingenuity in bacteria and highlights how microbes deploy unconventional strategies to survive viral attack. It also points to a vast reservoir of uncharacterized biology within microbial “dark matter,” where many fundamental mechanisms likely remain undiscovered.
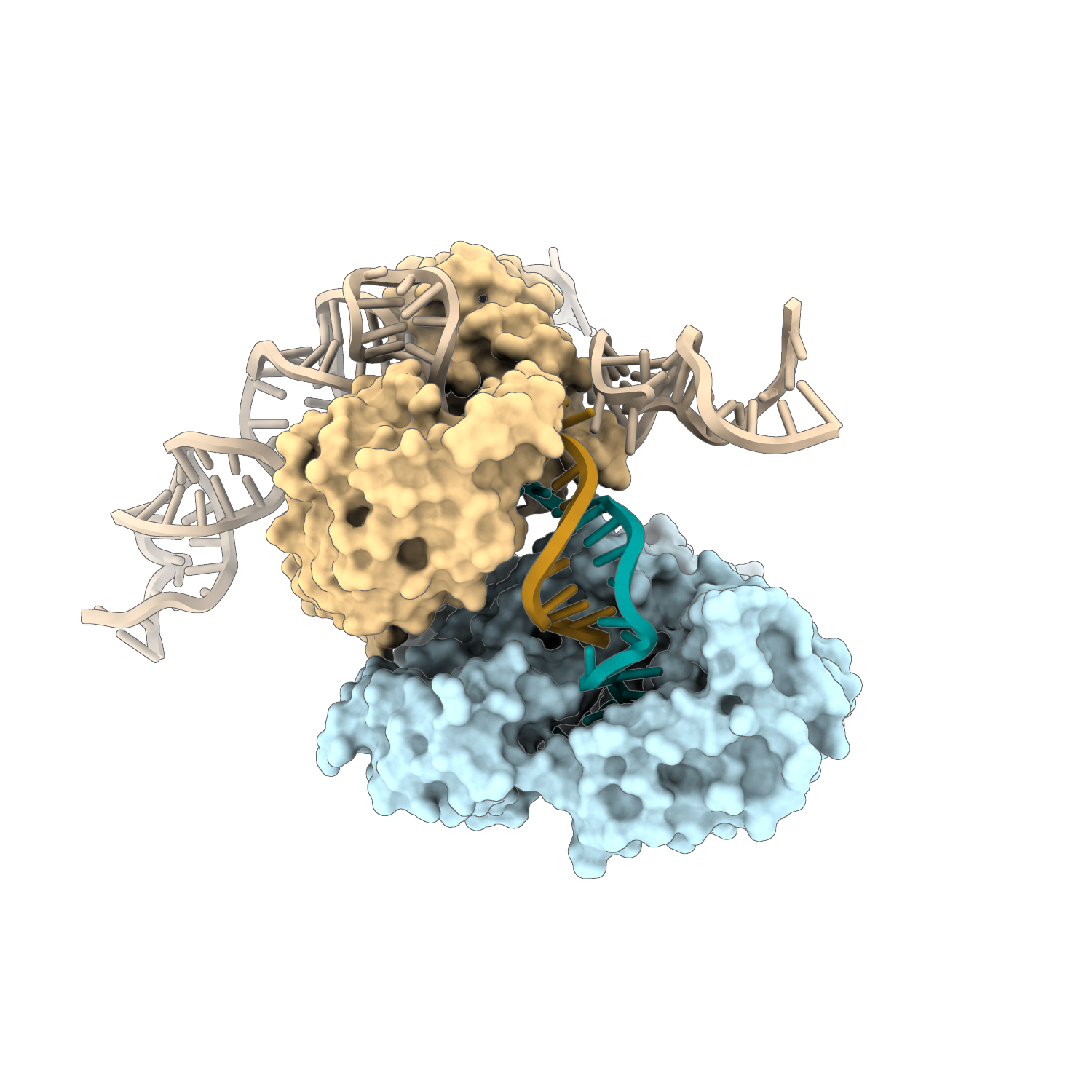
In the DRT3 bacterial defense system, paired strands of DNA (orange and cyan) are synthesized by two enzymes: one (yellow) uses an RNA template (beige) to guide DNA assembly, while a second, unusual enzyme (light blue) uses its own amino acids to build the complementary sequence.
Picture credit: Hyunbin Lee
Read the full article here. Paid subscription to Science required.


